UMSA
Untargeted Mass-Spectrometry Analysis
UMSA is an untargeted mass-spectrometry analysis service from RECETOX (Research Centre for Toxic Compounds in the Environment at Masaryk University) in the Czech Republic. The service is evolving to a key component of the emerging EIRENE ESFRI. By means of the integration in EOSC, uniform access to data and computing resources are provided, scaling the service to the target European-wide user community. Typically, mass spectrometry is done in a targeted way to confirm or disprove the presence of a specific compound in a sample. On the contrary, we aim at processing data to correlating the whole spectra (ie. all the present compounds) with other data (social, medical, other sample analyses, etc.) to work with more complex hypotheses of environmental impacts on human health.
The data are unrecoverable, original samples cannot be re-acquired, therefore long-term data storage (even decades) is required, together with appropriate data curation. Tracking provenance of the secondary (derived) datasets (what was the exact process of generating them from the original source data), is fairly critical, as the results may differ dramatically with different settings.
The current release provides a Galaxy workflow based on re-factored tools originating from Emory university (apLCMS and xMSAnnotator), which detect peaks in the input spectra and matches those to metabolite and pathway databases. Extending the workflow with auto-tuning peak picking parameters (based on the original xMSAnalyzer tool) and false-positive filtering by retention time prediction is in progress.
Architecture
The service is deployed as a virtual cluster using an Infrastructure Manager RADL recipe. The cluster consists of a single head node, running Galaxy frontend and the Slurm server, and an arbitrary number of Slurm worker nodes. Data is shared among the nodes over NFS.
The deployment of the head node registers a configurable dynamic DNS name (umsa.dyn.cerit-sc.cz currently) to point to its assigned IP address. The well known service endpoint (umsa.cerit-sc.cz) is expected to be an alias pointing to the dynamic one. In the next step a Let’s encrypt certificate is acquired to allow smooth https connection.
The RADL recipes also require simple provider-specific customization (cloud network names and base image identifiers in particular).
Authentication of the end users is managed by ELIXIR AAI via its corresponding Galaxy module. Configuration of Elixir AAI is the only manual step in the service deployment; the policy of introducing a new service to Elixir AAI requires human approval and exchange of secrets which cannot be automated so far. In the current version, the service can be accessed bona-fide — all users who pass ELIXIR AAI authentication are allowed. However, we plan a trivial registration procedure (using the ELIXIR tools) to restrict the access.
The payload of the service are several tools in Galaxy. In order to keep strict control on the complex software dependencies, we wrap all the software to Docker containers; Galaxy and Slurm are configured to execute them in this way only. The tools themselves are installed from standard Galaxy toolshed to follow common procedures.
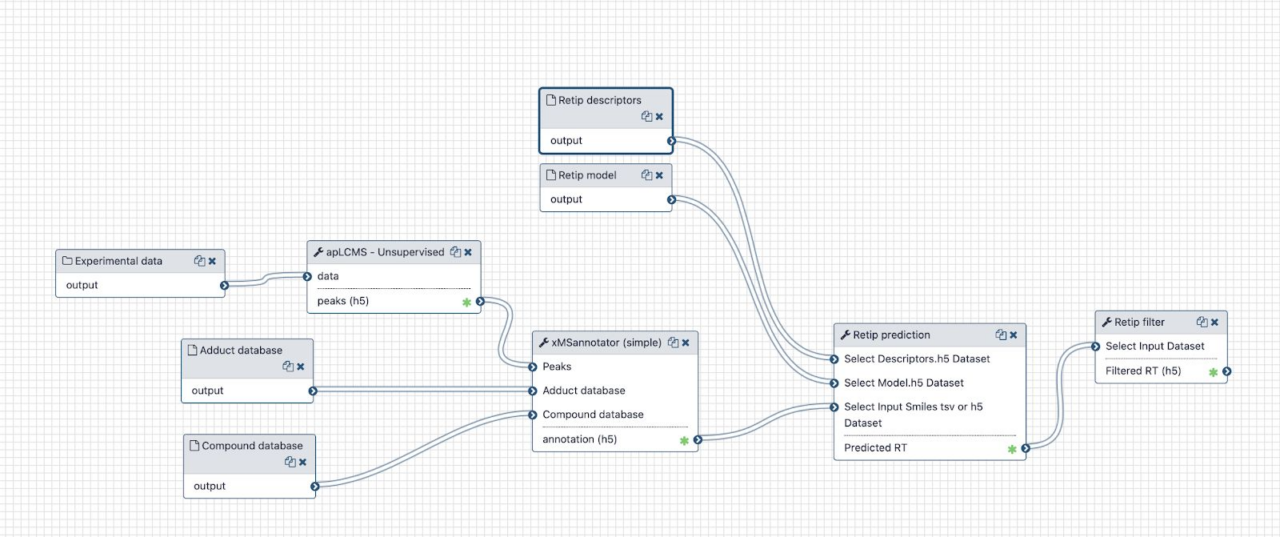
The diagram in figure shows the essential mass-spec data processing workflow. Besides the user input (.mzML spectra files) the workflow needs to query a metabolite database. In order to achieve reproducibility, timestamped snapshots of the online databases are used. They are downloaded and processed only from time to time, and the snapshots are available to all service users. Most of the intermediate files passed between the tools are some kind of tabular data, using the Apache feather format consistently.
EOSC Services
UMSA leverages the following EOSC services
- Infrastructure Manager (IM) and Elastic Compute Clusters in the Cloud (EC3): except from minor provider-specific customization the service is deployed with a generic RADL recipe submitted to IM, and IM takes care of its lifecycle.
- EOSC computing resources: the deployment relies on cloud computing resources; due to non-trivial computational demand of the workflow, “HPC” flavors of cloud virtual machines (with no CPU overprovisioning) are prefered. The production service runs at the CESNET/Masaryk University cloud site, we managed to deploy at CESGA successfully as well.
- EGI CheckIn: used to authenticate to IM as well as to deploy the VMs to the cloud sites. The service is currently using the “catch all” eosc-synergy.eu virtual organization.
Service Endpoint
The principal service endpoint is https://umsa.cerit-sc.cz/. The user interface is standard Galaxy with minimalistic visual branding. Login with Elixir AAI credential is required as described above.
The individual tools are available in PeakPicking and Annotation sections, both exist in simple and advanced forms to address the needs of different user experience levels. The in-line documentation provides extensive description of the meaning of numerous input parameters, as well as appropriate references to web pages with further documentation of the tools, and the essential journal papers.
Figure shows a typical input form of a simple workflow connecting the tools together.
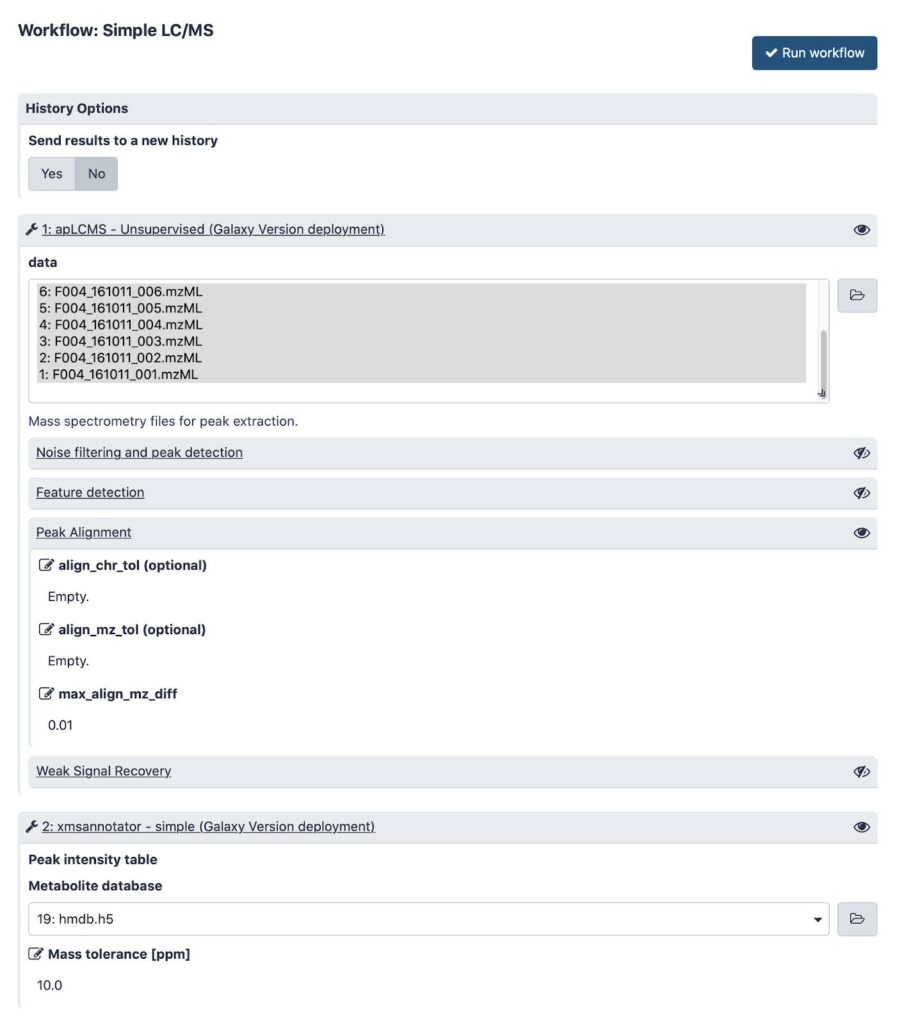
In this figure shows execution of this workflow in progress, generating its output files in Galaxy history.
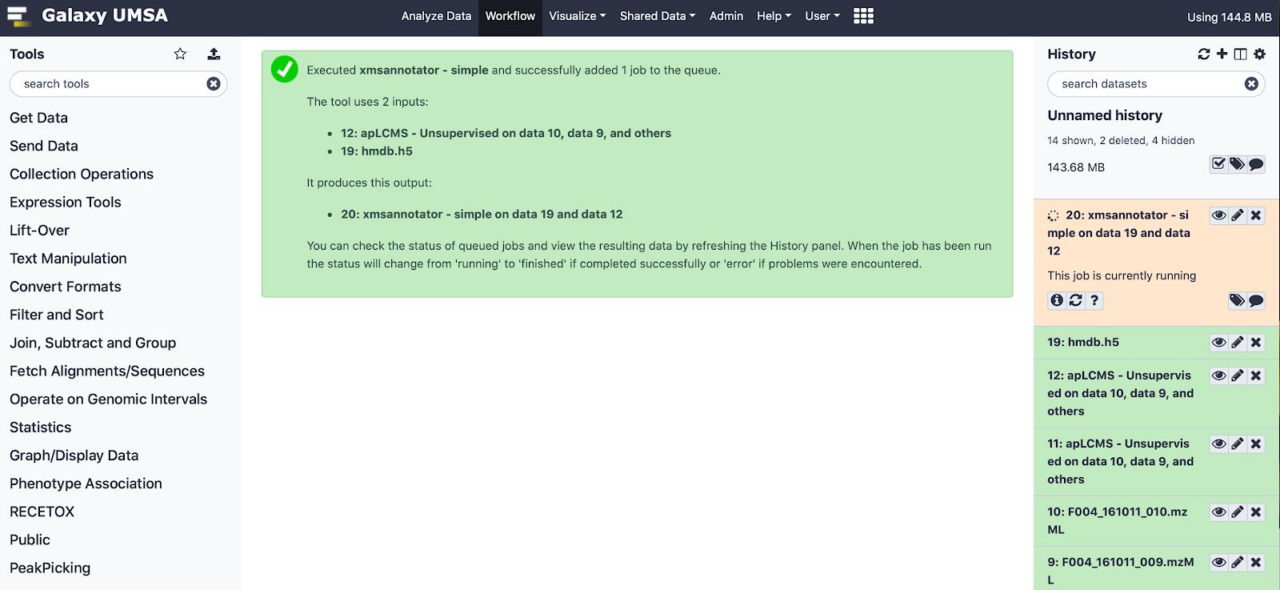
(apLCMS) has already finished, annotation is running.